- Home
- News & Updates
- Bringing AI-guided exploration to multiomics analysis
-
Illumina Connected Multiomics
-
Product updates
-
News
- 02/20/2026
Bringing AI-guided exploration to multiomics analysis
With the v1.1 release of Illumina Connected Multiomics, we’re adding AI‑assisted suggestions, our first step toward guided workflow construction.
AI is changing how scientists explore multiomic data and how insights are discovered. Instead of stitching together results across tools and modalities, researchers can now use AI‑guided exploration to surface meaningful biological signals earlier and more interactively, with far less dependence on deep bioinformatics expertise. This shift turns multiomics from a complex, specialist‑driven workflow into a guided discovery experience that helps teams move from data to insight faster and with greater confidence.
Illumina Connected Multiomics was built to make multiomic exploration practical at scale. With the v1.1 release, Connected Multiomics introduces AI‑assisted capabilities designed for one of the hardest parts of exploratory analysis: deciding what to do next. This AI‑powered guidance helps researchers navigate complex, integrated transcriptomic, epigenomic, proteomic, and genomic data efficiently, without sacrificing transparency or control.
A Platform for End-to-End Multiomics Analysis
Connected Multiomics organizes analyses as a visual task graph, where each step: QC, normalization, dimensionality reduction, clustering, or downstream interpretation, is represented as a node. This approach makes it easy to track provenance and reproducibility, branch analyses to test alternatives, and combine multiple data types in a single workflow.
The platform integrates seamlessly with DRAGEN™ for high-performance upstream processing and Correlation Engine for biological context and validation, supporting both routine analyses and open-ended exploration for users with varying levels of computational expertise.
Why AI-assisted workflows matter for exploratory science
In exploratory science, workflows are rarely fixed. Researchers iterate, branch, and refine analyses based on intermediate results. Across thousands of real-world workflows, consistent patterns emerge, with certain steps tending to follow others depending on data type and analytical context.
The new AI capabilities in Connected Multiomics surface these patterns and present them to the user for consideration. Instead of prescribing a static “best” workflow, the system provides context-aware suggestions that reflect how similar analyses are typically constructed in practice.
AI suggestions, full user control
In the v1.1 launch, Connected Multiomics will introduce AI-powered workflow suggestions as an optional, opt-out feature available to selected users.
- Suggestions are generated based on the current task
- Proposed steps appear as suggestions
- Each suggestion includes a confidence score and a brief rationale
- Users can accept, modify, or ignore suggestions at any point; when a suggestion is accepted, users manually configure parameters before running the analysis
This design keeps scientists firmly in control while reducing friction in workflow construction.
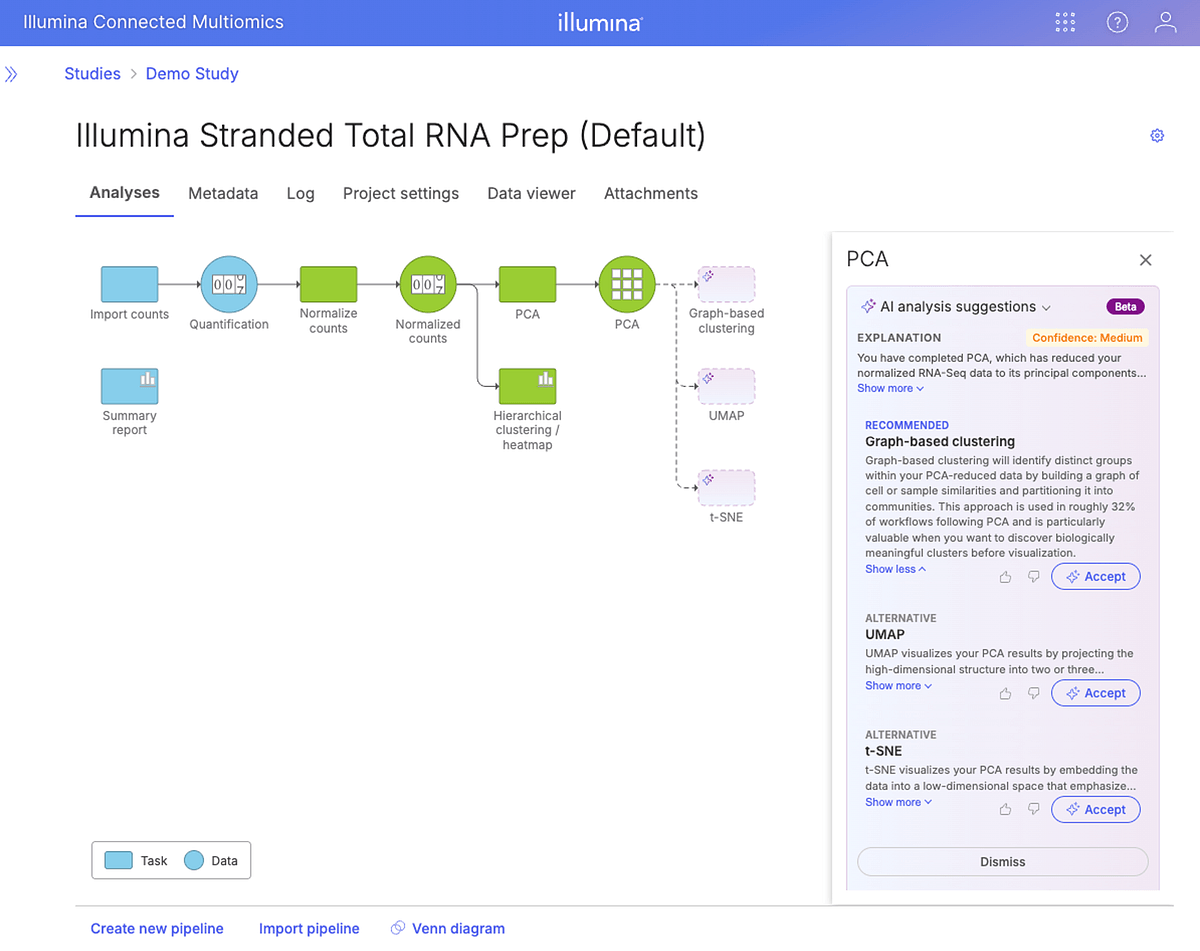
Conceptual comparison of a manually constructed multiomics task graph versus a task graph augmented with AI-proposed next steps. Suggested nodes are visually distinguished and can be selectively accepted or declined.
Evidence-based recommendations, not a black box
Under the hood, workflow suggestions are generated using a hybrid approach:
- Fast statistical prediction identifies likely next steps based on patterns learned from thousands of historical workflows and Illumina expert-created workflows that are weighted higher in the model.
- Natural-language explanations translate that statistical evidence into concise, domain-aware rationales.
Importantly, the AI does not execute analyses automatically. It proposes, explains, and defers final decisions to the researcher, preserving reproducibility and scientific judgment.
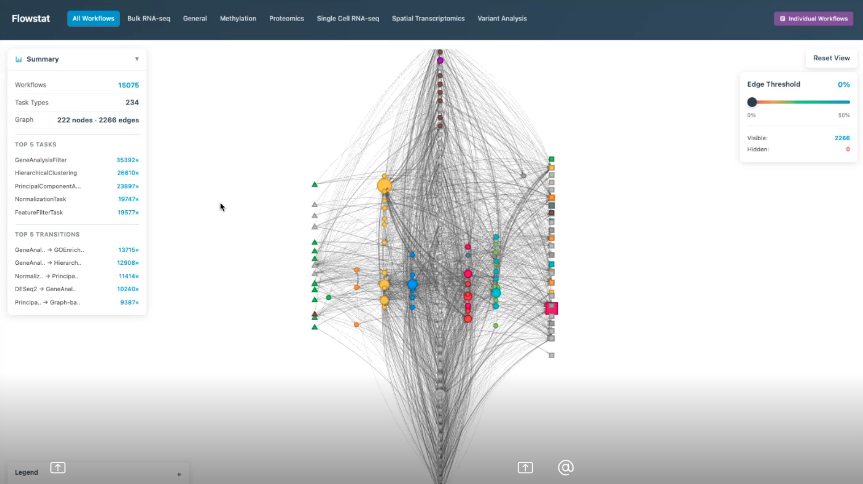
Task‑transition graphs derived from thousands of historical workflows, used as training data for Connected Multiomics AI‑powered workflow suggestions.
Designed for both learning and efficiency
For researchers new to Connected Multiomics or a particular modality, AI suggestions provide guardrails that help establish sensible analytical baselines. For experts, they function as an accelerator, reducing setup time for routine steps and surfacing alternative paths worth considering.
As multiomics analyses expand to include more data types and samples, predefining optimal workflows becomes impractical. Context-aware AI offers a scalable way to support exploration without constraining creativity.
Looking ahead
The v1.1 release marks the first integration of AI-guided workflow construction in Connected Multiomics. Future directions include richer multi-step suggestions, deeper multi-sample awareness, and more interactive ways to interrogate why specific analytical paths are recommended.
The goal is straightforward: less time assembling workflows, more time interpreting biology.
Ready to learn more or experience it for yourself?
Visit the product page | Watch demo videos | Request a free trial
---
For Research Use Only. Not for use in diagnostic procedures.
M-GL-04113