- Home
- News & Updates
- Enabling Cancer Interpretation At Scale For The Genomics England 100K Genomics Project
-
News
- 03/19/2019
Enabling Cancer Interpretation At Scale For The Genomics England 100K Genomics Project
Perspectives on training and on-boarding users of the Genomics England Cancer Program
By Jawahar Swaminathan, Ph.D., Program Manager - Population Genomics (aided by Keira Cheetham, Ph.D., Staff Bioinformatics Scientist)
Illumina and Genomics England announced the Bioinformatics and Clinical Interpretation partnership (BCIP) in February 2016 with the aim: “develop a platform and knowledge base that can be used to improve and automate genome interpretation.” As part of this collaboration Illumina developed a customized version of BaseSpace™ Variant Interpreter (BSVI)1 for cancer and rare disease, including various backend services to allow integration between the Genomics England case dispatch pipeline and Illumina systems. What followed was a rigorous schedule of meetings between Genomics England and Illumina (read as long hours, late nights, lots of coffee and many meetings at Genomics England HQ in London!) leading to development of essential features for cancer interpretation.
In June 2017, following multiple rounds of user acceptance testing and concordance checks, BSVI was adopted by Genomics England as the default interpretation solution. Illumina then began the process of on-boarding various users at the 13 Genomic Medicine Centres (GMC), the recruiting hubs for various regions of England by organizing training sessions on the use of the software with particular focus on the unique way data entered and left the system. This article is a look back on these activities and how they are helping in the development of genome interpretation software that meets the diverse needs of the Genomics England end users.
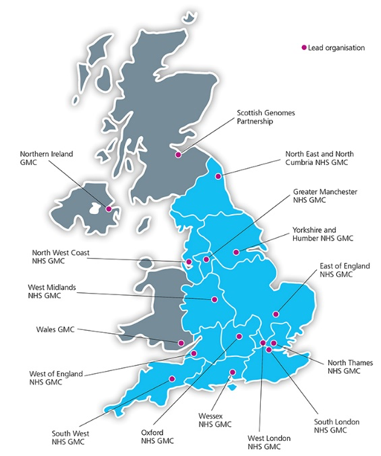
The GMC training sessions
Over the course of 2018, we carried out training and outreach activities across most of the GMCs. The GMCs are the recruitment hubs for the Genomics England 100,000 Genomes Project and comprise of multiple hospitals centered around a geographical area that has the necessary expertise. All training activities were organized by the Genomics England Cancer Interpretation team and were also attended by a representative from Genomics England.
Some humorous takeaways:
- Long hours on an early morning packed train from Cambridge (where we are situated) to our destination city, including a hurriedly eaten lunch at a busy Costa Coffee (yes almost every hospital in the UK has one of these) at the hospital before the training! Throw in the occasional aborted visit due to an alarmingly growing windscreen crack on a rental car or boarding the wrong train and you have the makings of a long and interesting day.
- Every NHS hospital looks the same. The usual 1960s concrete exterior, the same typeface on the signs and the same warren of corridors to the Clinical Genetics department
- Working out how to use the different display equipment in different hospitals before attempting to figure out internet connectivity on the slow and ageing hospital computer systems.
- Hot chocolate or a burrito on the return leg at the local train station as a treat for a job done well
- Never work with children, animals, or live demos. Although we always got the live demo to work!
All training activities were conducted by my colleague Keira Cheetham and I and involved a mix of presentations, live demos using cases specific to the GMC followed by hands-on instructions on how to use the software and send results back to Genomics England for reporting. The training was also an opportunity for us to talk about the science around interpreting cancer genomes and how Illumina is facilitating greater insights into cancers with whole genome sequencing (WGS).
This was also a great opportunity to see how the BCIP tools were used by GMC users and any feedback (both good and bad) were gratefully received. We also spoke about upcoming features in these sessions. Attendance at these events varied from 2-10 users per GMC and the venues ranged from really tight spaces (sometimes with windows!) to large meeting rooms and everything in between. However, what was consistent throughout was the motivation and dedication of the NHS staff in delivering the best possible care to their patients recruited into the Genomics England 100,000 Genomes Project cancer program.
Illumina continues to work with Genomics England to extend its BCIP tools for Rare Disease interpretation and this offering will soon be available for user acceptance testing and following that, could be used in Genomics England’s suite of clinical interpretation systems. In the meantime, the UK NHS has announced the commissioning of WGS for rare disease and cancer, to be offered throughout the health system. The outreach activities of 2018 carried out by Keira and I for cancer will keep in us good stead for the next round of training for rare disease.
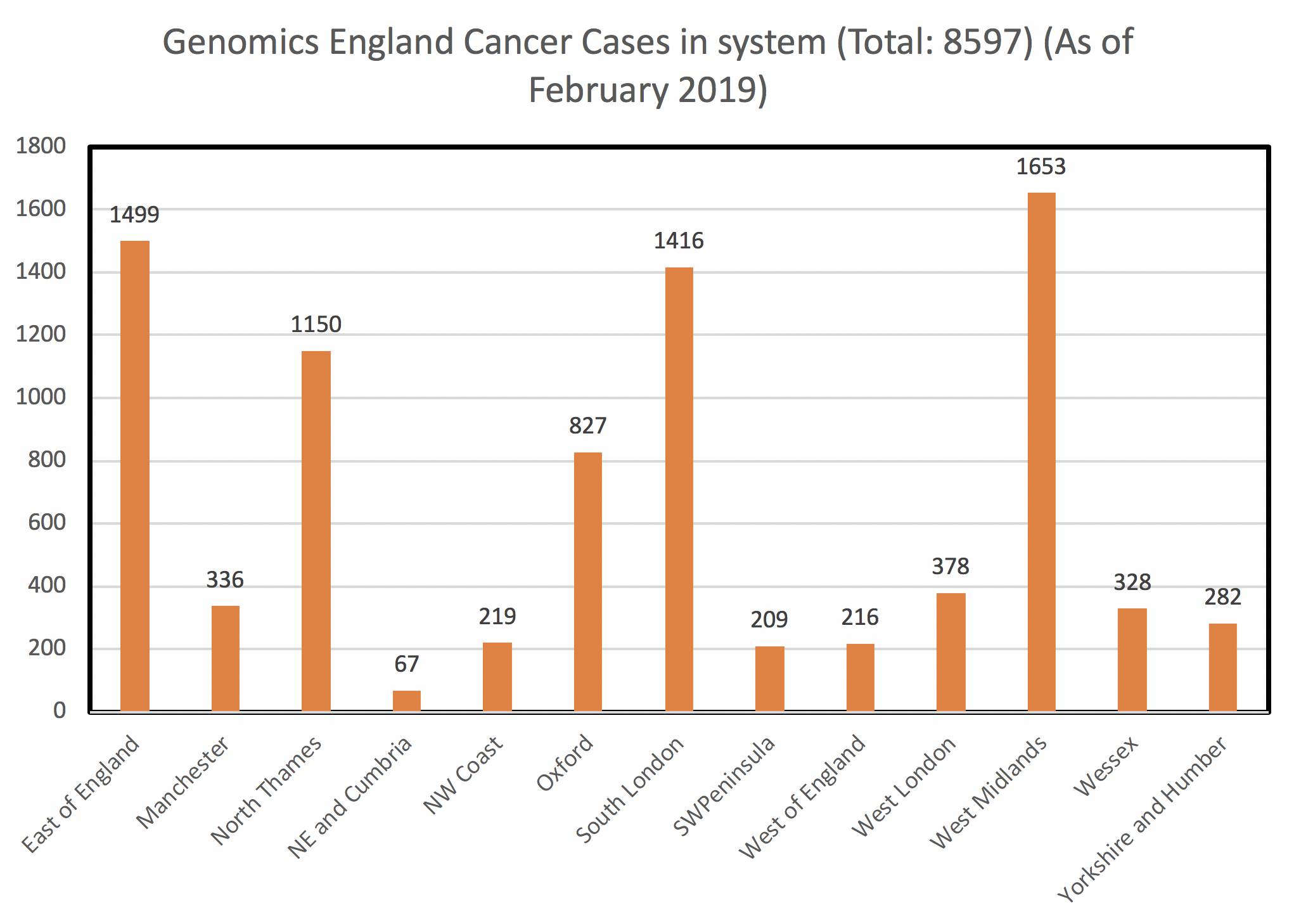
The Genomics England Cancer Outreach Program by numbers
- ~76 GMC users across 11 GMCs trained
- ~ 34 hours of training imparted
- ~4000 miles travelled (all by British Rail barring Belfast Northern Ireland)
- The version of BSVI co-developed with Genomics England as part of the BCIP contains extensive customizations for their use cases and is not openly accessible to the public. Please contact your Illumina sales representative for guidance on how to use the publicly available version of BSVI.