- Home
- News & Updates
- Accelerate omics pattern discovery with sophisticated multiomic analysis workflows leveraging Illumina’s Correlation Engine
-
Correlation Engine
-
Product updates
- 05/18/2023
Accelerate omics pattern discovery with sophisticated multiomic analysis workflows leveraging Illumina’s Correlation Engine
The field of multiomics is rapidly expanding, as researchers seek to understand the complex relationships between genes, proteins, metabolites, and other biomolecules. To help researchers unlock new insights and accelerate research, Illumina’s Correlation Engine provides a curated omics knowledge base of biological interactions that is driven by the vast amount of multiomic public data and publications, along with the comprehensive analytic correlation tools built-in.
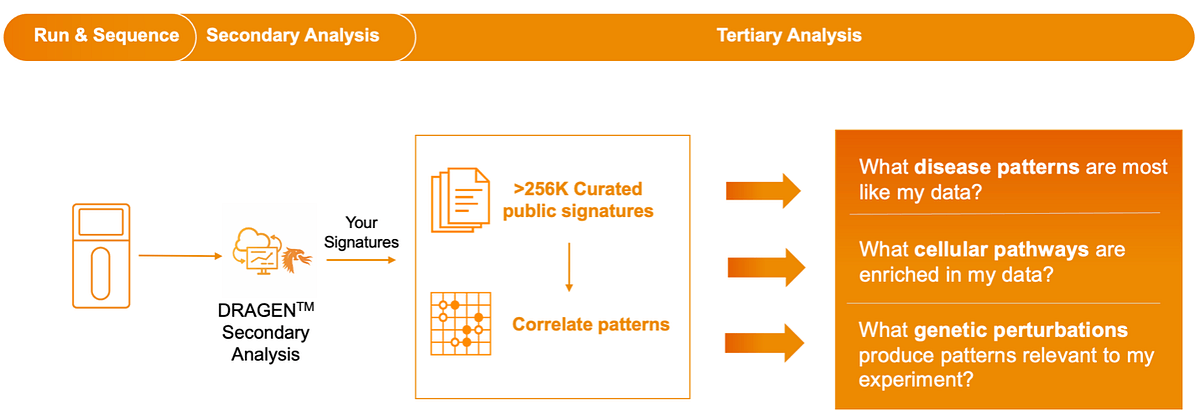
The interactive data analysis environment in Correlation Engine helps researchers validate and refine results and compare signatures from their own experiments with results from a large, curated repository of open-access and controlled-access public datasets (Figure 1). Correlation Engine's knowledge base indexes across the full mulitomics stack allowing users to measure one analyte, like the transcriptome, and see potential impact on another analyte, like the proteome, from others’ experiments. Users can further explore with a comprehensive suite of applications for determining biological context for use cases like biomarker discovery, drug repurposing, and drug safety and toxicogenomics.
With Correlation Engine’s latest release, we are excited to introduce a meta-analysis API (Application Programmable Interface) to facilitate programmatic access to the meta-analysis functionality. The Correlation Engine Meta-Analysis App lets users gather private and public genomic signatures of any of the supported datatypes (Figure 2), including the recently introduced ATAC-Seq, and any of the supported species (Figure 3). Researchers also have access to a vast number of insightful scientific tools including the Body Atlas, Disease Atlas, Pharmaco Atlas, Knockdown Atlas, Pathway Enrichment, and many more.
Supported Datatypes
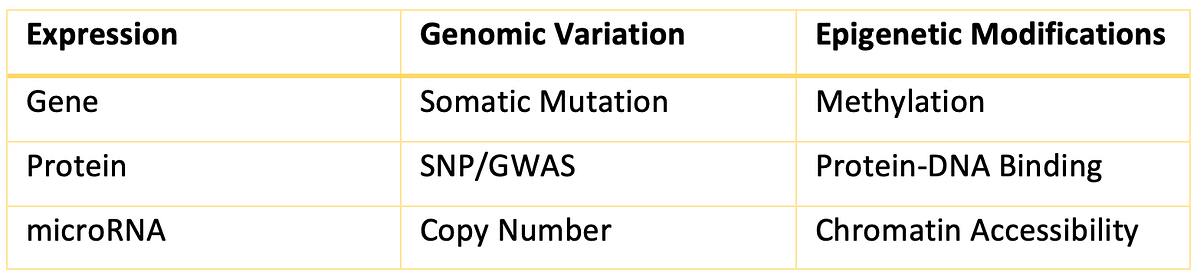

Supported Species

Multiomics is a compelling and increasing vital approach to understanding the etiology of disease and the impacts of chemicals and environmental exposures. In a recent publication from the United States Environmental Protection Agency, the University of Kansas Medical Center and Zunyi Medical college, the investigators used ChIP-Seq, RNA expression and microRNA expression data along with the resources of Correlation Engine to understand how environmental arsenic exposure led to epigenetic changes in the livers of developing embryos of mice that in turn led to phenotypic manifestations as adults (Liu et al., 2020). They identified genes impacted by differential methylation and miRNA expression and leveraged the Pathway Enrichment application in Correlation Engine to identify the top 50 altered changes of canonical pathways.
Researchers can also do in-depth genomic analyses by enabling integrations between Illumina Connected Analytics and Correlation Engine. An Illumina bundle of Rshiny applications provide tools for the analysis of bulk and single-cell RNA-Seq data with push-button import into Correlation Engine. With future plans to integrate more ICA tools with Correlation Engine, researchers can customize solutions for their specific analysis needs.
Learn More
To learn more about Correlation Engine, visit our website, and for an in-depth summary of all the latest Correlation Engine features, view our release notes.
Learn more on integrating multiomic methods into your research with Illumina solutions here.