- Home
- News & Updates
- Demo Data on Illumina® BaseSpace™ Sequence Hub
-
BaseSpace™ Sequence Hub
-
News
- 06/17/2021
Demo Data on Illumina® BaseSpace™ Sequence Hub
Today, we’re sharing the second in a series of posts that will shed more light on new run data publicly available on BaseSpace™ Sequence Hub (BSSH).
Prospective instrument and flow cell customers may wonder what they can expect from our latest sequencing platform, flow cell, or library preparation products. In this post, we cover how to use Illumina® BaseSpaceTM Sequence Hub (BSSH) to view and understand what typical sequencing runs on our platforms can look like. Any user with a free Illumina® account can log into BSSH to access and view these runs.
Illumina® BaseSpaceTM Sequence Hub contains a section called Demo Data (figure 1). This section provides test runs and their associated project data sets that you can import to your account. This helps users familiarize themselves with BaseSpaceTM Sequence Hub, and review typical data sets for a variety of applications from different sequencers and flow cells. These data sets are free and do not count against BSSH storage limits.
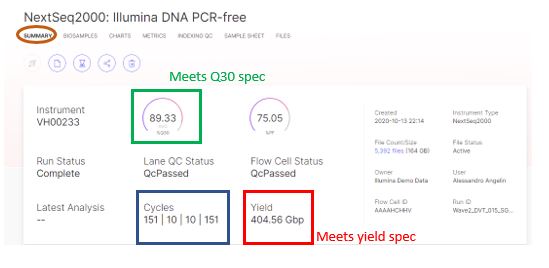
Figure 1: Screenshot of BaseSpace Sequence Hub homepage
To illustrate how to use the Demo Data tab, let us imagine that our lab recently acquired a NextSeq™ 2000 and we want to see what type of data quality to expect for samples prepared with Illumina DNA PCR-Free.
1/ Find data of interest
Data are organized by sequencing instruments, and they are listed in different categories and research areas such as Cancer Research or microbial research (figure 2):

Figure 2: demo data page showing publicly available sequencing runs, organized in various categories and research areas.
2/ Access run of interest
Select a dataset of interest, and then expand the arrow to access the Run and Project data. Run data will contain sequencing metrics and data from the flow cell that was sequenced. Project data will contain any secondary analysis of samples that have been demultiplexed from the Run data.
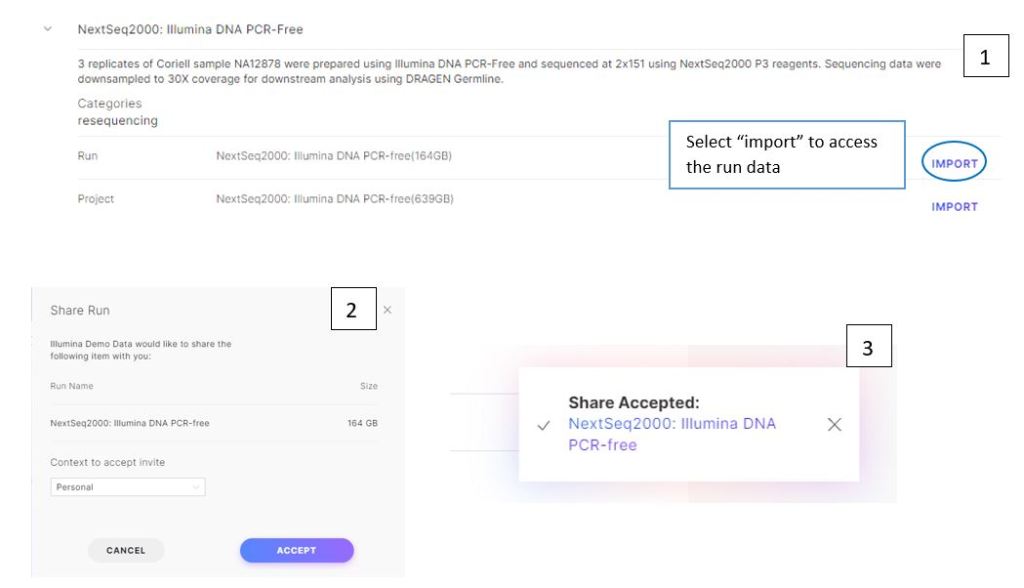
Figure 3: access run of interest. Click “Import” (1), then “Accept” (2) to access the runs. Once the share has been accepted, click the run name link to view the run data (3).
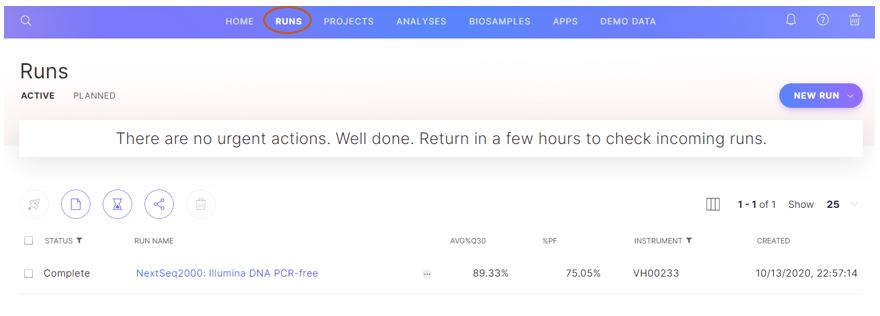
Figure 4: The demo run data can then be found under the “runs” tab for easy access in the future.
Now as we install and use the new NextSeqTM 2000 in our lab, we can know what a typical sequencing run will look like, and have something to compare against if we observe any anomalies. For more details on viewing sequencing run metrics on BSSH, view our [recent blog post on the topic](link).
3/ Access sequencing data of interest (or “project”)
Similar to a run, we will import the project (containing the secondary analysis data).
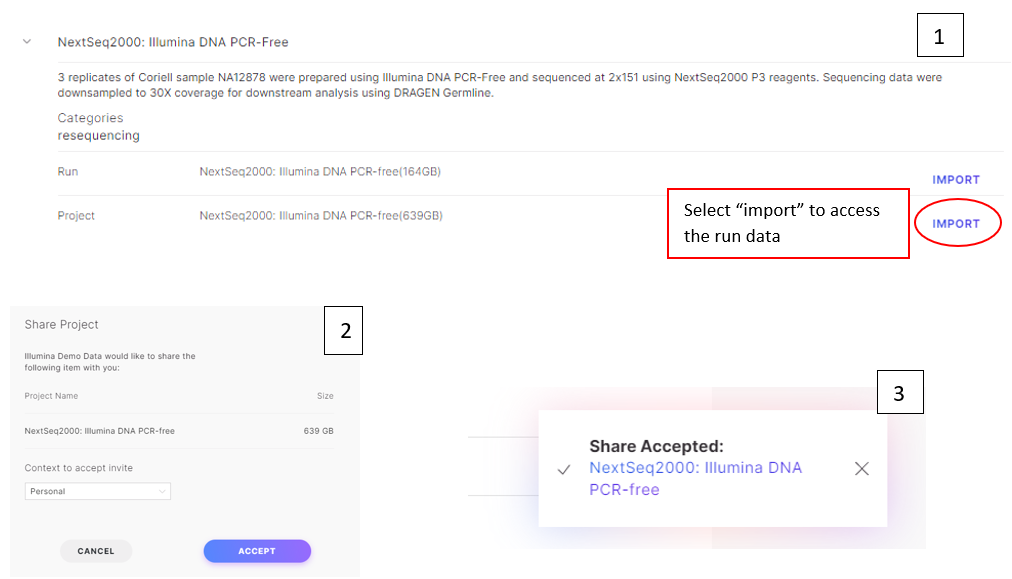
Figure 5: access data of interest (or “Project”). Click “Import” (1), then “Accept” (2) to access the projects. Once the share has been accepted, click the project name link to view the run data (3).
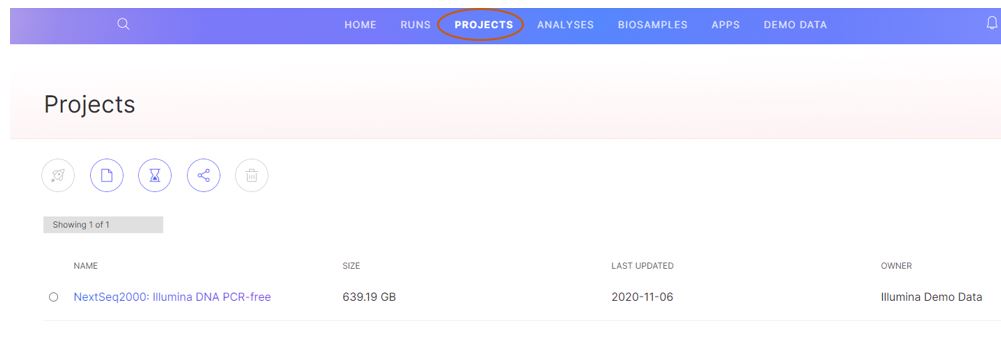
Figure 6: The demo project data can then be found under the “projects” tab for easy access in the future.
4/ Reviewing data analysis within the imported project folder
Within the Project folder that was imported from the Demo Data tab, there will be one entry for each of the analyses that were performed on the sequencing run data. In our example, there are DRAGEN Germline analyses for each Illumina® DNA PCR-Free sample that was sequenced on NextSeqTM 2000, followed by analysis using the Variant Call Assessment Tool (VCAT) to look at the resulting variant calls across all samples.
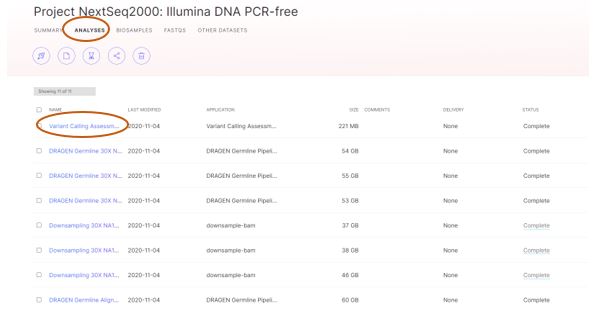
Figure 7: example of analyses data, for Illumina® DNA PCR-Free libraries run on the NextSeqTM 2000, and analyzed with DRAGEN Germline.
Looking at secondary analysis on the demo data page is free. Note that launching analysis with your own data using a BaseSpace app will require a subscription.
To learn more about BaseSpace Sequence Hub, contact your Illumina Representative.
Additional resources
BSSH product page: https://www.illumina.com/products/by-type/informatics-products/basespace-sequence-hub.html
BSSH help page: https://help.basespace.illumina.com/articles
Other posts from the blog series: new run data publicly available on BaseSpace™ Sequence Hub (BSSH).
- https://developer.illumina.com/news-updates/basespace-sequence-hub-does-my-sequencing-run-look-good
- https://developer.illumina.com/news-updates/10x-genomics-chromium-single-cell-multiome-atac-gene-expression-data-now-available-on-illumina-basespace-sequence-hub
Products mentioned here are for research use only, and are not for diagnostic use.